CNValidatron, automated CNVs validation in R using a CNN
Convolutional Neural Network trained on a vast collection of human validated CNVs, capable of automating the visual inspection of CNV calls from SNP arrays data. 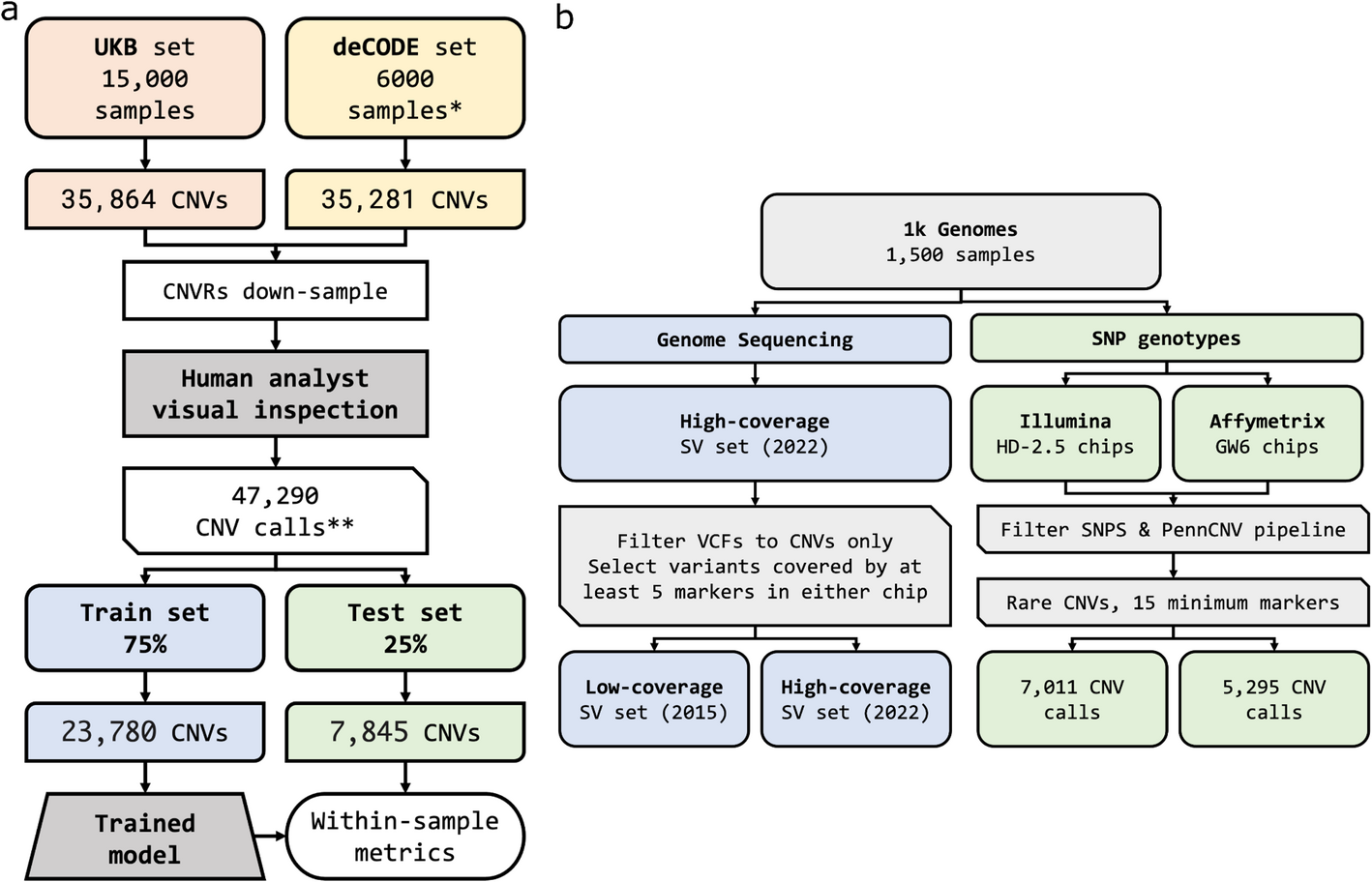
Convolutional Neural Network trained on a vast collection of human validated CNVs, capable of automating the visual inspection of CNV calls from SNP arrays data. 
Published in Current Protocols, 2022
Modern protocol to detct and process CNVs in large SNP genotype datasets, with a focus on biobank-scale resources. Open access!
Published in The Lancet Psychiatry, 2023
Large population-based study of recurrent sex chromosome aneuploidies and risk of psychiatric disorders in the iPSYCH cohort.
Published in JAMA Psychiatry, 2024
Large population-based study of recurrent copy number variants and risk of psychiatric disorders in the iPSYCH cohort.
Published in npj Genomic Medicine, 2024
Large population-based association study of exonic deletions at the NRXN1 locusand their risk of neurodevelopmental disorders. Open access!
Published in BMC Bioinformatics, 2025
This paper presents a novel approach for validating copy number variant calls using computer vision techniques. Open access!
Published:
I gave a short presentation at EurBioc 2024 on the on the development and application of the CNN model that would later be published in BMC Bioinformatics. It’s the same presentation I gave at NSHG-PM in Throndheim that year. You can find the slides of the presentation here.
Published:
In the context of the NSHG-PM 2024 conference, Andres and I were invited to hold a worshop on the topic of “AI, Data Science & Genomics Applications”. The course material is available on GitHub.
Published:
I gave a short presentation at NSHG-PM 2024 on the on the development and application of the CNN model that would later be published in BMC Bioinformatics. It’s the same presentation I gave at EurBioc in Oxford that year. You can find the slides of the presentation here.